Ultra-Fast 1H COSY
It cannot be disputed that the introduction of routine
2D NMR spectroscopy in the 1980's revolutionized the way in which NMR measurements are made. Now, with literally thousands of 2D methods available, the quantity of accessible information has dramatically increased. One cannot imagine a modern NMR lab without a 2D NMR toolbox. One of the main drawbacks to traditional 2D NMR spectroscopy has always been the time required to collect the data. Data collection can take anywhere from a few minutes to tens of hours. Many 1D FIDs (typically more than 128) must be acquired as a function of evolution time to construct the 2D data matrix. The measurement of each of these signals may require multiple scans as a result of necessary phase cycling between which a relaxation delay must be employed. Once all of the data have been collected, each of the FID's is Fourier transformed followed by a second Fourier transform with respect to the evolution time. Typical data collection and processing are illustrated
here. The introduction of pulsed field gradients for coherence selection has reduced the time required to collect 2D spectra by reducing or eliminating the need for phase cycling however, one still has to collect many FID's as a function of evolution time. Even when multiple scans are not required for sensitivity, data collection can take minutes to hours.
Ultra-fast 2D measurements, employing an entirely different method of data collection, were introduced in 2002 and subsequently improved. In this method, z-field gradients combined with linearly swept chirp pulses are used to phase encode spins linearly along the z axis of the sample according to specific evolution times. The dephasing depends on both the position along the z axis of the sample and the resonance frequency of each spin. After this encoding scheme is applied, each slice element of the sample has experienced a different evolution time as a function of its position in the sample. After a conventional mixing period dictated by the type of 2D measurement, the site specific, spatially phase encoded spins must be read. This is accomplished by applying a series of bipolar gradient pulse pairs while the receiver is collecting data. During each gradient pulse (lasting typically 250 ?sec) echos are collected. The position of each echo during a single gradient pulse is related in a one-to-one fashion to the frequency of each of the spins in the sample thus mimicking a mini NMR spectrum whose frequency axis is replaced by a linearly related time axis. The "spectra" collected during the negative gradient pulses are mirror images to those collected during the positive gradient pulses and must be reversed during data processing. A series of typically 128 bipolar gradients are applied with the receiver open thus all of the data are acquired in a single scan. Each "spectrum" collected is a function of the z slice position in the sample, which in turn is linearly related to the evolution time. The collection of "spectra" represent the ultra-fast domain and is Fourier transformed point by point as a function of evolution time (or z position). The entire data collection sequence takes approximately 100 msec.
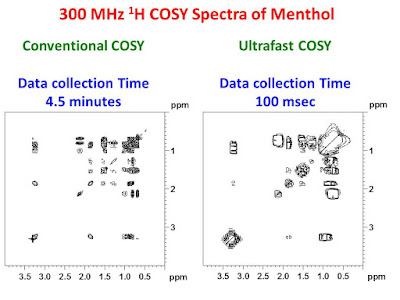
The left panel of the figure below shows a conventional 300 MHz gradient enhanced COSY-45 spectrum for a concentrated sample of menthol in CDCl3 collected in 4.5 minutes. The panel on the right shows a 300 MHz ultra-fast COSY spectrum of the same sample collected in only 100 msec - a time saving factor of 2700! Both spectra were collected on a Bruker AVANCE II 300 NMR spectrometer equipped with a standard BBOF probe. Both data sets were symmetrized. Although the ultra-fast data set has noticeably lower resolution and sensitivity, one can see that it is very similar to the conventional COSY.
There are, of course, a number of drawbacks to the ultra-fast scheme including low sensitivity, limited resolution and limited accessible spectral widths. Some of these drawbacks can be overcome with the use of cryoprobes and strong pulsed field gradients. Molecular diffusion over the course of the measurement may also cause problems. Despite the drawbacks however, the method is extremely well suited to time studies of chemical reactions where conventional 2D data collection would simply take too long.
The references below are a good place to start in order to find out more about this technique. There is also a very well documented setup procedure available on the
Bruker User Library, provided by
Patrick Giraudeau, including pulse sequences and processing scripts.
Annual Rev. Anal. Chem. 7, 129-161 (2014).
Mag. Res. Chem. 53, 986-994 (2015).
J. Am. Chem. Soc. 125, 920417 (2003).
J. Am. Chem. Soc. 125, 1234550 (2003).


Source:
University of Ottawa NMR Facility Blog



